
1
Orthomyxoviruses
اﻟﻤﺮﺣﻠﮫ اﻟﺜﺎﻟﺜﮫ /ﻓﺎﯾﺮوﺳﺎت
.م
د. اﻧﺘﻈﺎر ﻋﻼوي ﺟﻌﻔﺮ / ﻓﺮع اﻻﺣﯿﺎء اﻟﻤﺠﮭﺮﯾﮫ / ﻛﻠﯿﮫ اﻟﻄﺐ / ﺟﺎﻣﻌﮫ ذي ﻗﺎر
PhD. M.Sc. Microbiology
Introduction
These viruses are classified under two families
These are:
•
Orthomyxoviridae, consisting of influenza viruses.
•
Paramyxoviridae, consisting of parainfluenza, mumps, measles,
respiratory syncytial.
•
The genus Orthomyxovirus includes influenza viruses, the causative
agents of worldwide epidemics of influenza.
Influenza Viruses
Influenza viruses belong to the family of Orthomyxoviridae and are the causative
agents of influenza, a respiratory disease in humans (Flu)with well-defined
systemic symptoms that occurs in sporadic, epidemic, and pandemic forms.
Influenza A and B viruses cause substantial morbidity and mortality in humans
and a considerable financial burden worldwide, whereas influenza C viruses
cause sporadic outbreaks of mild respiratory disease, mainly in children.
N.B
v
Epidemic refers to an increase, often sudden, in the number of cases of a disease
above what is normally expected in a population within a geographic area
v
Pandemic refers to an epidemic that has spread over several countries or
continents, usually affecting a large number of people.
Properties of the Virus
Morphology
Influenza viruses are spherical or filamentous, enveloped particles 80–120 nm in
diameter.
It is composed of a characteristic segmented negative sense, single-stranded
RNA genome, a nucleocapsid, and an envelope (See-Fig).
Source: ViralZone:www.expasy.org/viralzone, SIB Swiss Institute of Bioinformatics)

2
The viral genome is a single-stranded antisense RNA. The genome consists of an
RNA-dependent RNA polymerase, which transcribes the negative-polarity
genome into mRNA.
• The RNA genome is segmented and consists of eight segments in
Influenza A & B and seven segments in influenza C viruses. These
segments code for different proteins, which are NS1, NS2, NP, M1, M2, M3,
HA, and NA.
The genome is present in a helically symmetric nucleocapsid surrounded by a
lipid envelope. The envelope has an inner membrane protein layer and an outer
lipid layer. The membrane proteins are known as matrix or M protein and are
composed of two components M1 and M2.
•
Two types of spikes or peplomers project from the envelope:
(a) The triangular hemagglutinin (HA) peplomers and
(b) The mushroom-shaped neuraminidase (NA) peplomers.
Classification and Nomenclature
Ø
Genus Influenza virus A contains human and animal strains of influenza
type A
Ø
Influenza virus B contains human strains of type B,
Ø
Influenza virus C contains influenza type C viruses of humans and swine.
Ø
Antigenic differences exhibited by two of the internal structural proteins,
the nucleocapsid (NP) and matrix (M) proteins, are used to divide
influenza viruses into types A, B, and C.
Ø
Antigenic differences exhibited by two of the internal structural proteins,
the nucleocapsid (NP) and matrix (M) proteins, are used to divide
influenza viruses into types A, B, and C.
Ø
Antigenic variations in the surface glycoproteins, HA and NA, are used to
subtype type A viruses
Ø
So far, 18 subtypes of HA (H1–H18) , 11 subtypes of NA (N1–N11), in
many different combinations, have been recovered from humans and
animals. e.g of current subtypes of Influenza A viruses H1N1 , H2N3
•
The standard nomenclature system for influenza virus isolates includes
the following information: type, host of origin, geographic origin, strain
number, and year of isolation.
•
Antigenic descriptions of the HA and the NA are given in parentheses for
type A. e.g
A/Hong Kong/03/68(H3N2)
Hemagglutinin (HA)
Ø
The HA protein of influenza virus binds virus particles to susceptible cells
and is the major antigen against which neutralizing (protective)
antibodies are directed.
Ø
Variability in HA is primarily responsible for the continual evolution of
new strains and subsequent influenza epidemics.

3
Ø
Hemagglutinin binds with the sialic acid cell receptor, and initiates the
infection in the host cell.
Ø
HA derives its name from its ability to agglutinate erythrocytes under
certain conditions.
Ø
The HA agglutinates certain RBC , which is inhibited by the neutralizing
Abs. This forms the basis of the hemagglutination inhibition test used in
the serodiagnosis of influenza.
Ø
Hemagglutinin has potency to undergo antigenic variations.
Neuraminidase (NA)
The NA is a glycoprotein and tetramer. It consists of 100 mushroom-shaped
spikes.
v
The NA functions at the end of the viral replication cycle. It cleaves the
neuraminic acid and to release progeny virions from the infected host
cells.
v
It is a sialidase enzyme that removes sialic acid from glycoconjugates.
v
NA facilitates release of virus particles from infected cell surfaces during
the budding process and helps prevent self-aggregation of virions by
removing sialic acid residues from viral glycoproteins.
v
It is possible that NA helps the virus negotiate through the mucin layer in
the respiratory tract to reach the target epithelial cells.
Antigenic variations
Antigenic variation is a unique feature of influenza virus.
The surface antigens HA and NA show variations and are primarily responsible
for antigenic variations exhibited by influenza viruses. The internal RNP antigen
and M protein are stable, hence do not contribute to the antigenic variations.
Antigenic variations are of two types: antigenic shift and antigenic drift.
Antigenic shift
²
Antigenic shift: Occurs due to major antigenic changes in HA or NA
antigens
²
Caused by replacement of the gene for HA by one coding for a completely
different amino acid sequence.
²
The antigenic shift is characterized by alteration of virtually all the
antigenic sites of the HA.
²
Demonstrated in type A influenza virus only.
²
Antigenic shift variants appear less frequently, about every 10 or 11
years.
•
Antigenic shift reflects drastic changes in the sequence of a viral surface
protein, caused by genetic reassortment between human, swine, and
avian influenza viruses.
•
The completely novel antigens that appear during antigenic shift are
acquired by genetic reassortment.
•
Influenza B and C viruses do not exhibit antigenic shift because few
related viruses exist in animals.
4
Antigenic drift
•
Minor antigenic changes are termed antigenic drift
•
Occurs due to minor antigenic changes in the HA or NA occurring at
frequent intervals.
•
Antigenic drift is caused by the accumulation of point mutations in the
gene, resulting in amino acid changes in the protein. Sequence changes
can alter antigenic sites on the molecule such that a virion can escape
recognition by the host’s immune system.
•
Antigenic drift variants occur very frequently, virtually every year.
Key Points
•
Influenza A virus shows maximum antigenic variations.
•
Influenza B virus does not undergo antigenic shift because influenza B
virus is the only human virus for which there is no animal source of new
RNA segments. However, influenza B virus undergoes antigenic drift.
•
Antigenic variation never occurs in type C influenza virus beacuse its lack
of NA.
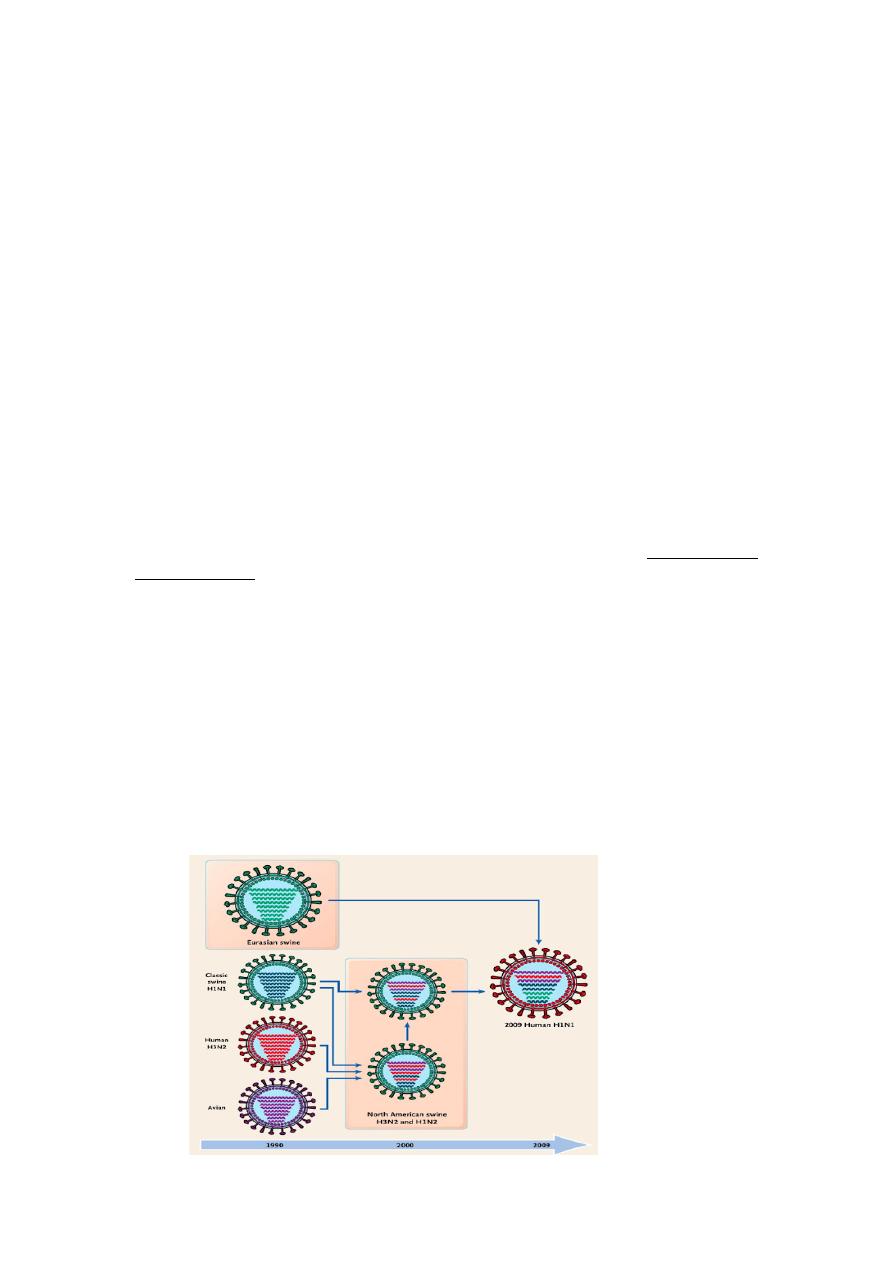
Gene reassortment
Because the influenza virus genome is segmented, genetic reassortment can
occur when a host cell is infected simultaneously with viruses of two different
parent strains. This process of genetic reassortment accounts for the periodic
appearance of the novel types of influenza A strains that cause influenza
pandemics.
Influenza viruses of animals, such as aquatic birds, chickens, swine, and horses
show high host specificity. These animal viruses are the source of the RNA
segments that encode the antigenic shift variants that cause epidemics among
humans. For example, if a person is infected simultaneously by an avian and
human influenza strains, then it is possible that a genetic reassortment could
occur in infected cells in humans. The reassortment could lead to emergence of a
new human influenza A virus, the progeny of which will contain a mixture of
genome segments from the two strains (e.g., a new variant of human influenza A
virus bearing the avian virus HA).
5
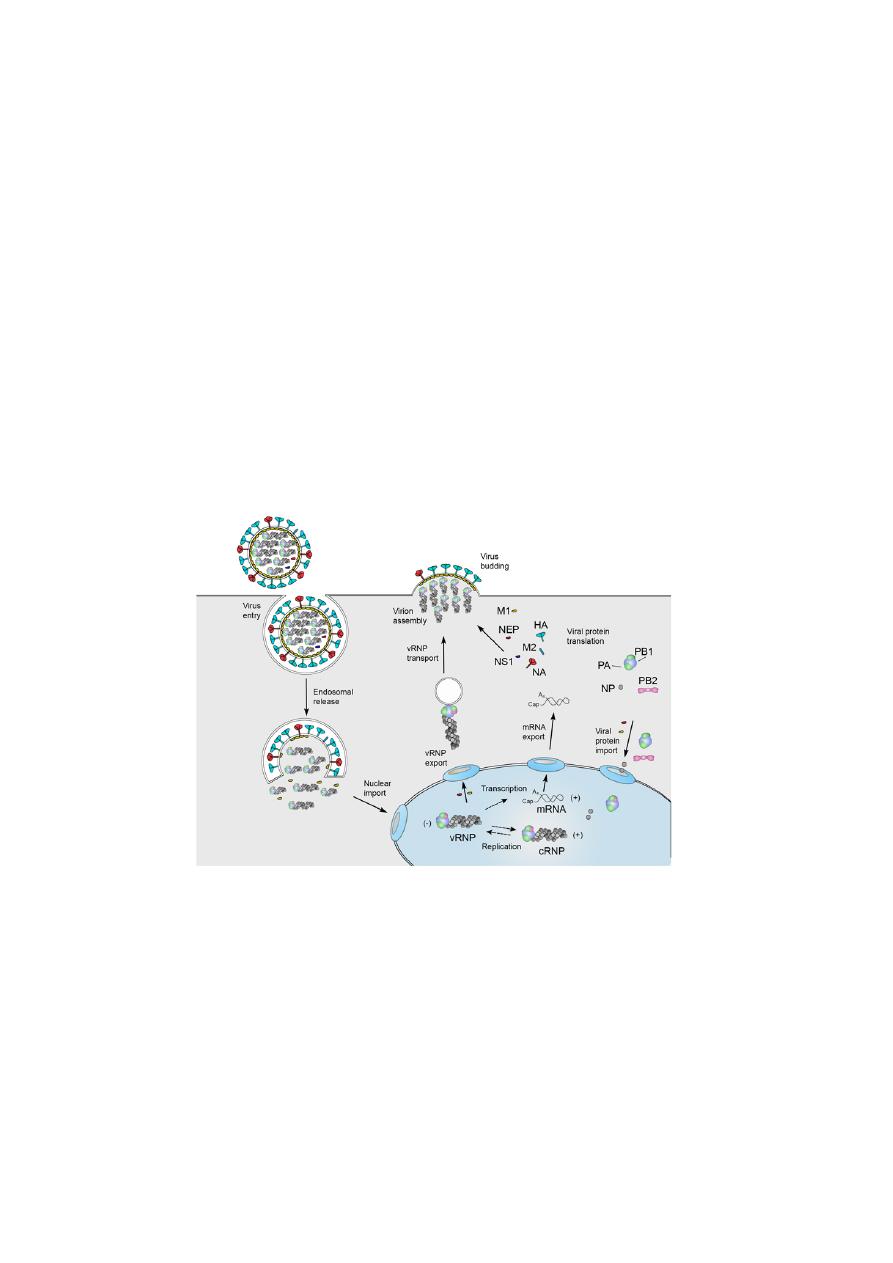
REPLICATION of Influenza virus
•
Viral infection initiates with the binding of a virion to cell surface
receptors containing sialic acid, followed by the endocytosis of the virion.
•
After fusion of the viral and endosomal membranes, the viral
ribonucleoproteins (vRNPs) are released into the cytoplasm and then
transported into the nucleus.
•
In the nucleus the viral RNA polymerase transcribes the vRNA segments
into mRNAs.
•
Viral mRNA is exported to the cytoplasm for translation by cellular
mechanisms. The viral RNA polymerase also performs replication of
vRNA by copying it into complementary RNA (cRNA), which serves as a
template for the production of more vRNA. Newly synthesised viral
polymerase and nucleoprotein are imported into the nucleus and bind to
cRNA and vRNA to assemble vRNPs and cRNPs, respectively.
•
Following nuclear export, progeny vRNPs are transported across the
cytoplasm on recycling to the cell membrane, where assembly of progeny
virions takes place.
•
Mature virions are released by budding.
Infleunza A virus replication cycle.
Source:
Aartjan J.W. te Velthuis and Ervin Fodor, 2017.
Pathogenesis and Immunity
Influenza virus is transmitted from person to person primarily in droplets
released by sneezing and coughing.
Inhaled influenza viruses reach lower respiratory tract, tracheobronchial tree,
the primary site of the disease. They attach to sialic acid receptors on epithelial
cells by HA present on the viral envelope. Relatively few viruses are needed to
infect lower respiratory tract than the upper respiratory tract. Neuraminidase of
the viral envelope may act on the N -acetyl neuraminic acid residues in mucus to
produce liquefaction.

6
Infection of mucosal cells results in cellular destruction and desquamation of the
superficial mucosa. The resulting edema and mononuclear cell infiltration of the
involved areas are accompanied by symptoms including nonproductive cough,
sore throat, and nasal discharge. Although the cough may be striking, the most
prominent symptoms of influenza are systemic: fever, muscle aches, and general
prostration. The virus remains localized to the respiratory tract; hence viremia
does not occur. In an uncomplicated case, virus can be recovered from
respiratory secretions for 3–8 days.
Clinical Syndrome
Incubation period is short (1–3 days). The classic influenza syndrome is a febrile
illness of sudden onset, characterized by tracheitis and marked myalgias.
Headache, chills, fever, malaise, myalgias, anorexia, and sore throat appear
suddenly. The body temperature rapidly rises to (38.3–40.0°C) and respiratory
symptoms ensue. Nonproductive cough is characteristic. Sneezing, rhinorrhea,
and nasal obstruction are common.
Patients may also report photophobia, nausea, vomiting, diarrhea, and
abdominal pain. They appear acutely ill and are usually coughing. Minimal to
moderate nasal obstruction, nasal discharge, and pharyngeal inflammation may
be present.
Complications
1-Secondary bacterial infections: Life-threatening influenza is often caused by
secondary bacterial infections with staphylococci, pneumococci, and
Haemophilus influenzae. Pneumonia may develop as a complication and may be
fatal, particularly in
(a) Elderly persons above 60 years with underlying chronic disease.
(b) In people chronic cardiorespiratory disease, renal disease, etc.)
(c) Pregnant women.
2-Reye’s syndrome is a noted complication of influenza B infection. The
condition is seen most commonly in young children and is associated with
degenerative changes in the brain, liver, and kidney.
3- Central nervous system complications: Guillain–Barre syndrome
characterized by encephalomyelitis and polyneuritis is a rare complication of
influenza virus infection.
Reservoir, source, and transmission of infection
Infected humans are the main reservoir of infections for influenza A virus.
Respiratory secretions of infected persons are the important source of infection.
The virus is excreted in respiratory secretions immediately before the onset of
illness and for 3–4 days thereafter. Wild aquatic birds are known reservoirs of
influenza A. They secrete the viruses in their feces, which contaminates ponds
and lakes. The virus is spread from person-to-person primarily by air-borne
respiratory droplets released during the acts of sneezing and coughing.
Influenza B virus only causes epidemics. Infection is from humans-to-humans.
No animal reservoir hosts are known.
7
Laboratory Diagnosis
During an epidemic of influenza, the clinical diagnosis can be made, but
definitive diagnosis depends on the laboratory methods.
Specimens:
•
Nasal or throat washings or sputum for viral antigen and viral RNA.
•
Throat gargles for isolation of viruses.
•
Serum for viral antibodies.
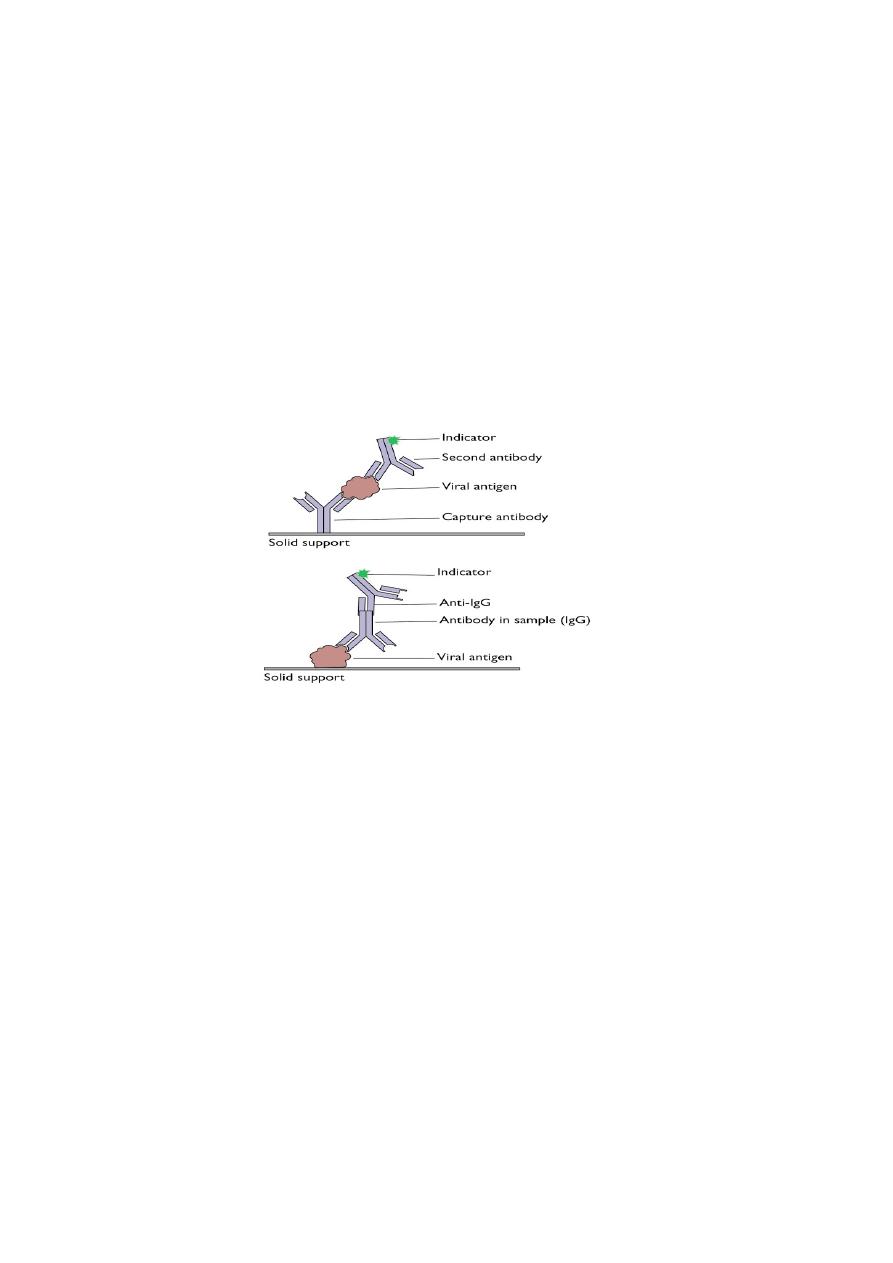
Direct antigen detection
Is made by demonstrating viral antigens directly on cells obtained from the
nasopharynx. Immunofluorescence (IF) or enzyme-linked immunosorbent assay
using specific monoclonal antibodies are used to detect viral antigen.
The results of the rapid tests are useful to start treatment with the NA inhibitors
within 48 hours of the onset of symptoms.
Detection of antigens by ELISA
Isolation of the virus
Throat gargles are the specimen of choice. The specimen is collected in saline
broth or a buffered salt solution and is sent immediately to the laboratory, or if
delayed is stored at 4°C.
The virus is isolated from the specimen by inoculation into embryonated eggs or
into certain cell cultures.
Treatment
Amantadine and Rimantadine are the specific antiviral agents available for
treatment of influenza. Tamiflu ((Oseltamivir phosphate)
Prevention
This is based on the following:
Immunoprophylaxis by vaccines: Influenza A subtypes H1N1 and H3N2 are most
common predominate human influenza viruses.
